Setting up an R workflow
Workflow
Why a clear data analysis workflow?
- Check analysis and track errors
- Share results with colleagues for stories or editing
- Send methodology to sources for bullet-proofing
- To easily adjust when presented with new data
- Easily switch between work environments (desktop and laptop)
- Scavenge and repurpose code in future projects
Constraints
- Workflow has to be platform agnostic
- Easy to deploy for yourself and others
- Free open source software
- Input has to be real raw data in whatever format it is (and wherever it is)
- But have a backup for when internet is not accessible
- Output has to work – whether html, PDF, or web app
- IDE agnostic (be able to run it from a command line without RStudio)
Four components
- Software
- R
- RStudio
- Git for version control
- Clear file organization
- One R script to pull it all together
- Hosting the html output internally or publicly with GitHub pages
Thanks
These are all things I picked up from browsing other presentations and repos.
Much thanks to Jenny Bryan and Joris Muller from whom I cobbled many of these ideas and practices from.
Also to BuzzFeed, FiveThirtyEight, ProPublica, Chicago Tribune, Los Angeles Times, and TrendCT.org
Use projects to organize
Do not dump your scripts into a folder
One folder per project
- RStudio project
- Git repo
- Can run parallel projects

Use portable file paths
DO NOT USE setwd()
Keep everything relative to your project directory and it will work on everyone who downloads your project repo folder.
Don’t run the lines of code below. This is just to show you how it would work in theory.
#install.packages("here")
library(here)
here("Test", "Folder", "text.txt")
##> [1] "/Users/IRE/Projects/NICAR/2018/workflow/Test/Folder/test.txt"
cat(readLines(here("Test", "Folder", "text.txt")))
##> You found the text file nested in these subdirectories!Files organization
At minimum
name_of_project
|--data
|--2017report.csv
|--2016report.pdf
|--summary2016_2017.csv
|--docs
|--01-analysis.Rmd
|--01-analysis.html
|--scripts
|--exploratory_analysis.R
|--name_of_project.Rproj
|--run_all.ROptimal
name_of_project
|--raw_data
|--WhateverData.xlsx
|--2017report.csv
|--2016report.pdf
|--output_data
|--summary2016_2017.csv
|--rmd
|--01-analysis.Rmd
|--docs
|--01-analysis.html
|--01-analysis.pdf
|--02-deeper.html
|--02-deeper.pdf
|--scripts
|--exploratory_analysis.R
|--pdf_scraper.R
|--name_of_project.Rproj
|--run_all.REverything below is for more advanced users but I’m putting it here for future reference.
Creating folder shortcut
folder_names <- c("raw_data", "output_data", "rmd", "docs", "scripts")
sapply(folder_names, dir.create)Organization principles
- Directory names are obvious to anyone looking
- Reports and the script files are not in the same directory
- Reports are sorted using 2-digit numbers. Tell your story clearly.
Source to the online data
Normal data file
if (!file.exists("data/bostonpayroll2013.csv")) {
dir.create("data", showWarnings = F)
download.file(
"https://website.com/data/bostonpayroll2013.csv",
"data/bostonpayroll2013.csv")
}
payroll <- read_csv("data/bostonpayroll2013.csv")Dealing with a zip file
if (!file.exists("data/employment/2016-12/FACTDATA_DEC2016.TXT")) {
dir.create("data", showWarnings = F)
temp <- tempfile()
download.file(
"https://website.com/data/bostonpayroll2013.zip",
temp)
unzip(temp, exdir="data", overwrite=T)
unlink(temp)
}
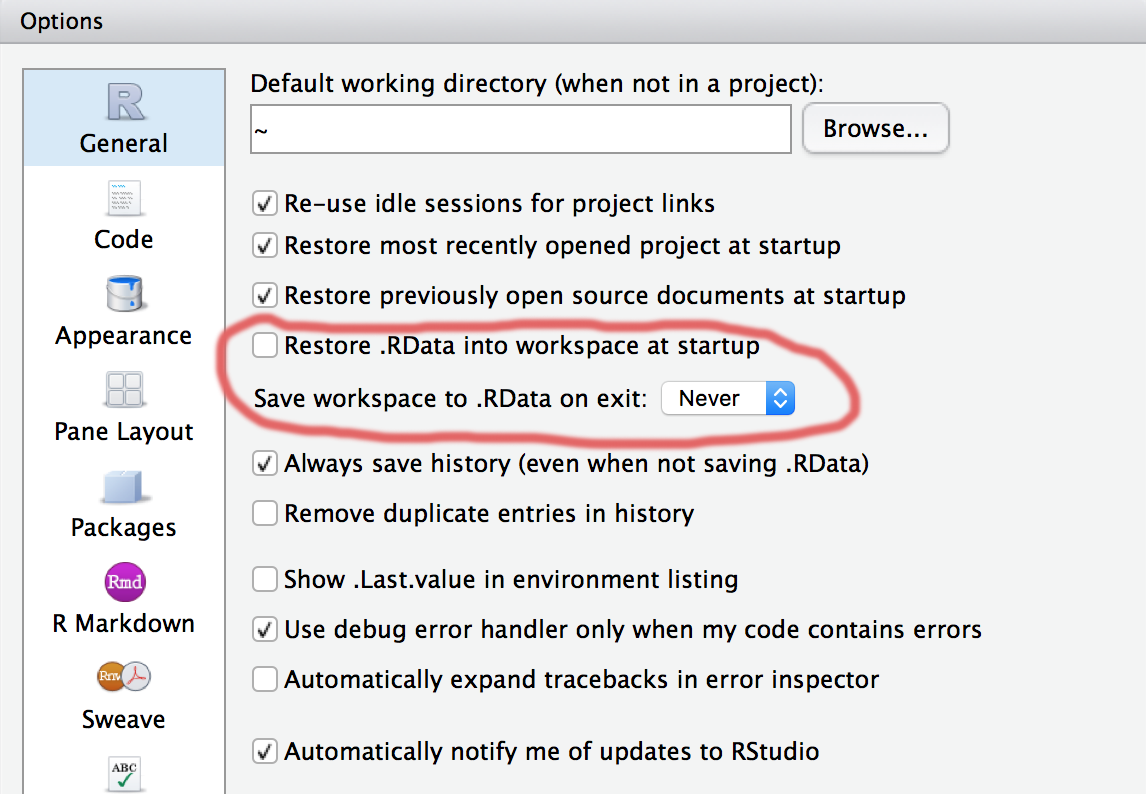
payroll <- read_csv("data/bostonpayroll2013.csv")Operate without a net
Never save work space to .RData on exiting RStudio and uncheck Restore .RData on start up.
This will make sure you’ve optimized your data ingesting and cleaning process and aren’t working with a misstep in your process.
© Copyright 2018, Andrew Ba Tran
Comment your code
Anything that appears on a line after
#will be treated as a comment. That means it will be ignored when the code in the script is executed.Use this to explain what the code does.
Get into this habit early. Future readers of the code will be grateful for the clear documentation you leave behind– including yourself months from now.