RMarkdown
By now you’ve inadvertently witnessed the different content that R Markdown can help generate:
- HTML documents that are in each chapter sub folder
- Notebooks in which you’ve run code chunks individually
- PDFs that you can print out to follow along physically with the course
- This entire learn.r-journalism.com website
R Markdown is R code + Markdown.

What is Markdown?
A very easy way to add formatting to plain text created by John Gruber (of Daring Fireball) as a simple way for non-programming types to write in an easy-to-read format that could be converted directly into HTML.
In the image below, the text on the left was interpreted as HTML on the right.

GitHub has a nice guide on Markdown.
R Markdown / .Rmd files
It’s a simple way to embed chunks of R code (or other languages like Python) in Markdown documents.
On one level, it allows you to present your analysis process and results in a format that doesn’t require R to run.
But the code is still there should a researcher or fellow journalist be compelled to reproduce or add to your work.
It’s a version of literate programming. By combining your R code with documentation, it makes your programming more robust, portable, and easier to maintain.
At the click of a button, or the type of a command, you can rerun the code in an R Markdown file to reproduce your work and export the results as a finished report.
R Markdown works thanks to the knitr package, which runs code embedded in Markdown, and Pandoc, which then converts Markdown into a bunch of different output formats, like Word, PDF, HTML, etc.
R Markdown supports dozens of static and dynamic output formats including
- HTML
- MS Word
- Beamer
- HTML5 slideshows
- Tufte-style handouts
- Books
- Dashboards
- Scientific articles (white pages)
- Entire websites
Formats useful to journalists
Produce slick-looking PDFs
Exporting your work into PDFs can be effective.
For this to work, be sure to get LaTex installed first.

IPython Notebooks
Notebooks are very popular.
This is how pandas and ipython notebooks render when uploaded to GitHub.
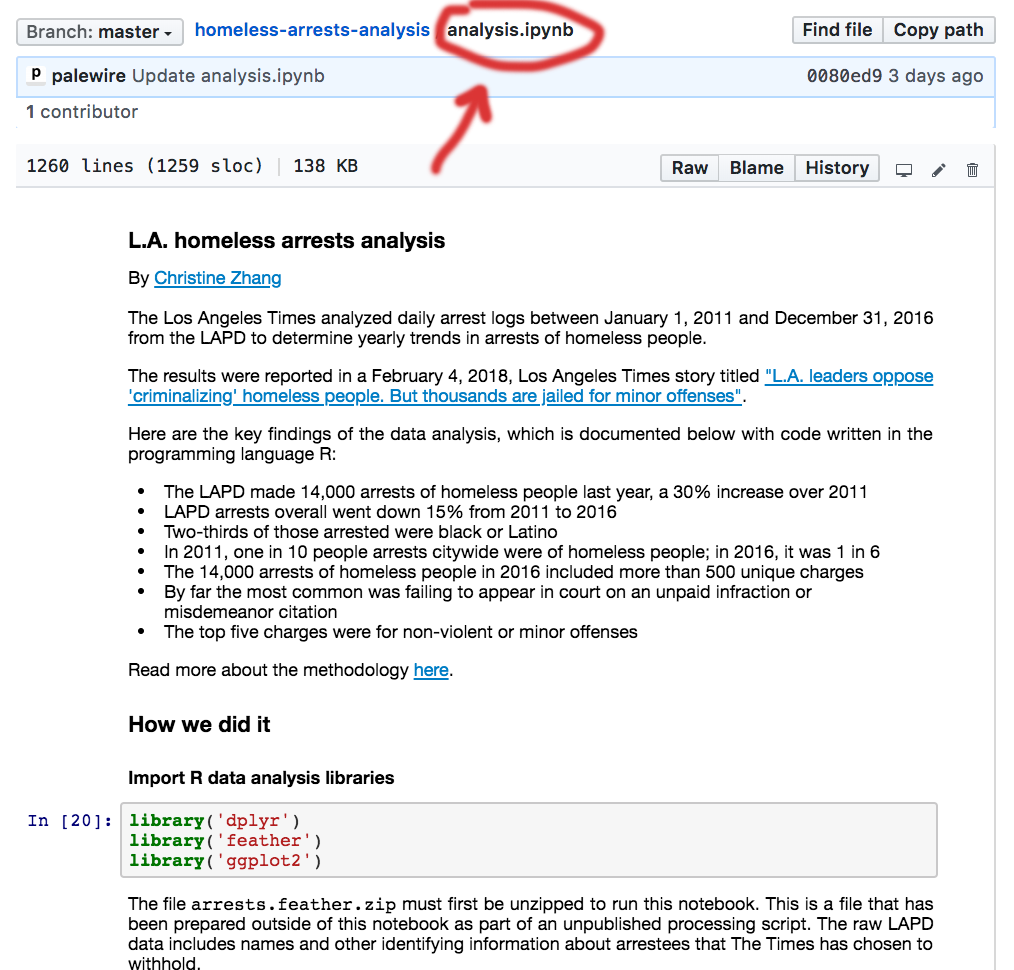
It renders well and matches the experience of someone coding in that environment.
.Rmd files rendered on Github
How does an R Markdown file look on GitHub?
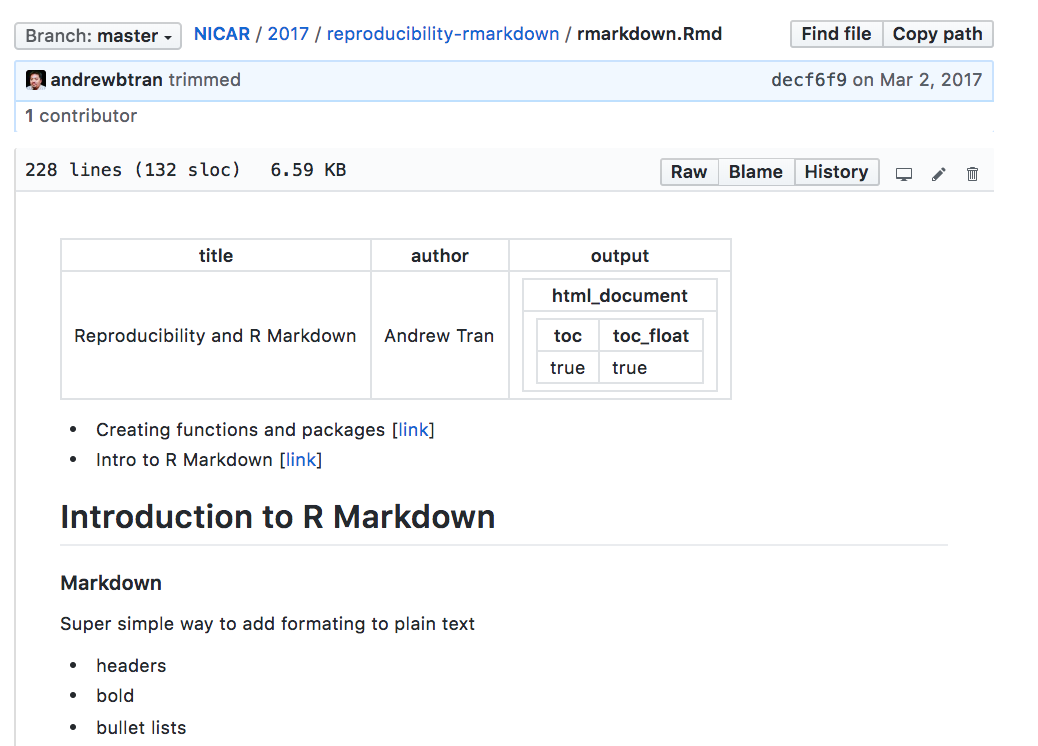
Not well.
Each .Rmd file has its own custom YAML section at the top. These are keywords that when combined with the right packages, lets knitr know how to output the .Rmd file. Like toc: toc_float creates an HTML file with a self-generated Table of Contents based on the header titles.
GitHub doesn’t have a way to interpret that so it creates that nested image above and doesn’t even try to make a table of contents.
That’s fine, though
Output .Rmd files to HTML
R Markdown will let you output as HTML, which you can still host on GitHub Pages (which we’ll go over later). You may have to include links to the actual page but doing so can be more effective than having a .Rmd file “render” in a GitHub repo.
Rendering in HTML lets you add CSS and JavaScript, allowing for the inclusion of content like the interactive table below.
library(DT)
library(dplyr)
library(readr)
payroll <- read_csv("../data/bostonpayroll2013.csv") %>%
select(NAME, TITLE, DEPARTMENT, REGULAR, OVERTIME) %>%
filter(row_number()<100)
datatable(payroll, extensions = 'Buttons', options = list(
dom = 'Bfrtip',
buttons = c('copy', 'csv', 'excel', 'pdf', 'print')
)
)This table was rendered with the DT package.
And we don’t have to create the code to display it.
It just worked because the folks who ported the DataTables jquery plugin over to R wanted to make it seamless.
Pass certain arguments to the function datatable() and you can include buttons that allow reporters to download your tables as CSVs.
Can you imagine the power of that?
You’ve transformed the data collected it and then instead of sending them a huge spreadsheet, you send them a link to your report in which they can filter things out and download the table that they’ve come up with themselves.
You can render other interactives, as well, like leaflet maps.
Links not attachments!
This is a very important development as a journalist.
You see, reporters sometimes aren’t very organized.
How often have you had someone email you asking if you could resend your summarized spreadsheet to them again?
This is what their downloads or desktop folder might look like. It’s easy to see why they might have lost track of it.

Send them links instead!
Host the HTML of your analysis
- On your internal work network or
- On Github pages (for free)
- Share with reporters and editors, let them explore your analysis further
- Let them download customized spreadsheets with buttons in the
DT(datatable) package - If your analysis gets updated, keep the file name then they only have to refresh their link
Repeat this process enough and maybe you can start learning how to create Shiny apps with R.
Creating R Markdown
Follow the instructions below.
1. Open a new .Rmd file
at File > New File > R Markdown.
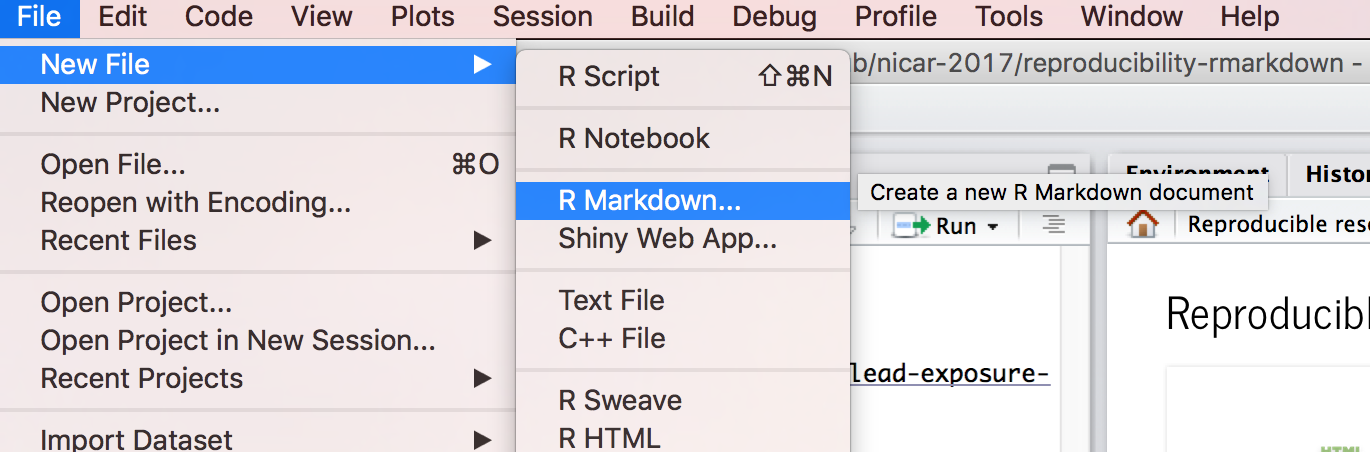
Title the R Markdown file and select HTML as the output for now.
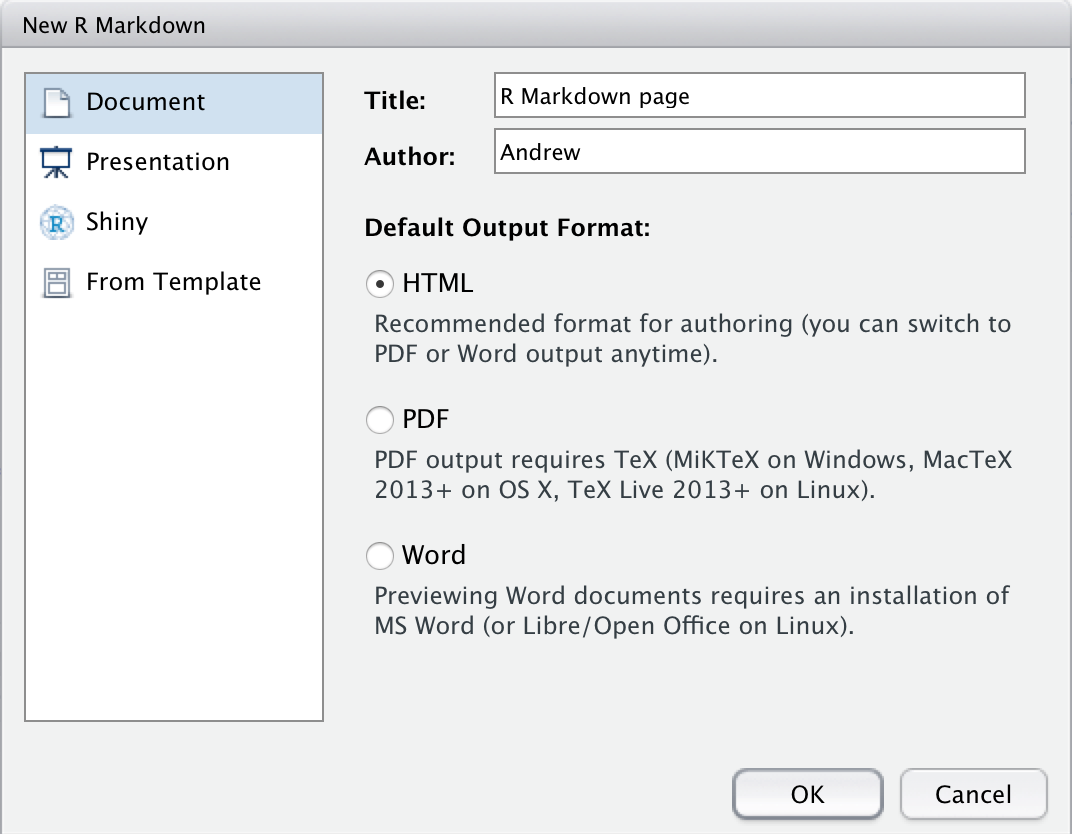
The default file created comes with code already included to give you a guidance on syntax and how it should be structured.
.Rmd structure
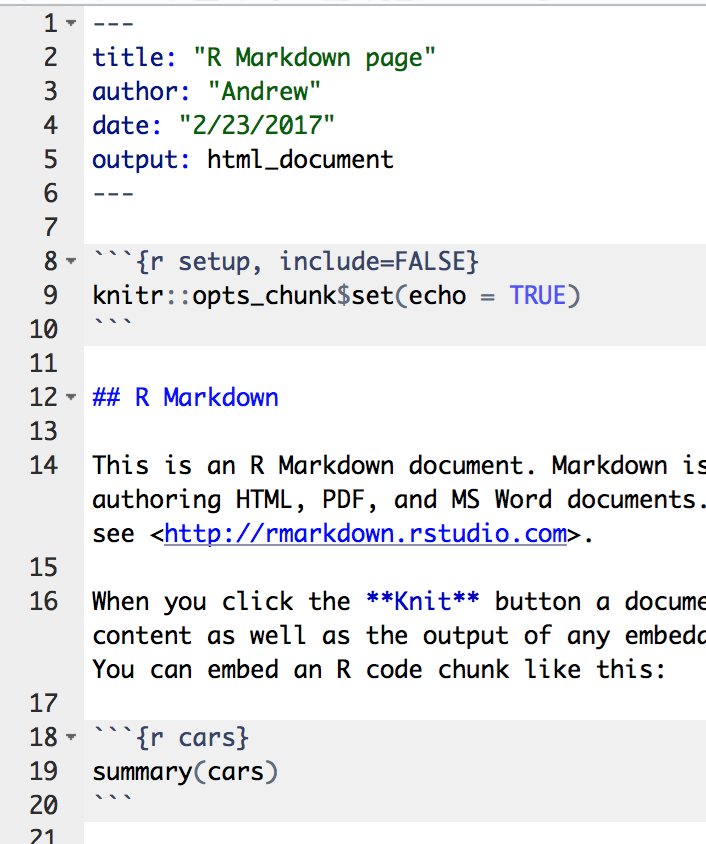
Every .Rmd file starts out the same at the top.
YAML HEADER
Optional section of render options written as key:value pairs.
- At start of file
- Between lines of — (3 dashes)
This is how R knows how to render your files. In the example above, the lineoutput: html_document means it will create an HTML file. If it was output: pdf then you’d get a PDF instead.
You can output more than one type at a time, if you want.
CODE CHUNKS
In the image above, the R code is in the gray area.
These are chunks of embedded R code. Each chunk:
- begins with ```{r} (the back tick key is to the left of 1)
- ends with ```
TEXT
The rest of it is just the narrative of markdown interspersed in between the R code chunks.
You can also use HTML code instead of Markdown if that’s what you’re more comfortable with.
2. Write document
Edit the default template by putting in your own code and text.
Intersperse the text with your code to tell a story.
2b. Label your chunks of code
It’ll be easier to navigate to in the future with the pull down menu at the bottom of the script window.
It also will be useful to narrow down where code errors might occur.

2c. Notebooks style
You can run individual chunks of code before generating the full report to see how it looks.
Click the green arrow next to each chunk.
Hopefully, you’ve already been doing this in our coding walkthroughs in class.

3. Knit document to create report
Use knit button (the yarn ball!) at the top left or type render() to create the output file(s).

3b. Check out the build log
Warnings and errors will appear in the console below.
Also measures progress by chunks, which is why it’s important to label them.

4. Preview output in RStudio window

5. Output file
You started with a .Rmd file and clicking knit HTML also generated a .html file.

It’s all there in your project folder now.
Next, we’ll go over more specific features in R Markdown files that will give you more control on how its presented.
© Copyright 2018, Andrew Ba Tran